Introduction to our research
Our research delves into the captivating world of enzyme mechanisms! Our primary goal is to unravel the intricate mysteries surrounding the functionality of enzymes. Have you ever pondered how these remarkable catalysts precisely identify and interact with their substrates? Or visualized the complex interplay between their structural components? Our research explores fundamental questions about how these biological marvels work. At the core of our investigations lies a fusion of cutting-edge methodologies: from pre-steady state kinetics to single molecule techniques, along with structural and biophysical approaches. Through these tools, we construct precise quantitative models, enabling a deeper comprehension of how enzymes operate within the cellular environment. Our current focus is on enzymes involved in three distinct and pivotal biological processes: DNA Repair & Recombination, mRNA recognition, and long-range electron transfer reactions in oxidoreductases. These areas serve as the frontier of our exploration, offering insights into critical cellular functions such as maintenance of genomic stability, RNA transcription & translation, and core chemistries such as nitrogen fixation and chlorophyll biosynthesis. In addition, mutations, often associated with diseases such as cancer, perturb function and deregulate enzyme function. Our work provides the mechanistic basis for why mutations result in disease. Our structural work aims to reveal suitable target sites for the development of small molecule-based chemotherapeutics. If the dissection of the inner workings of enzymes and unravelling their significance in cellular processes peek your interest, feel free to reach out or consider joining our quest for a profound understanding of these remarkable biological machines.
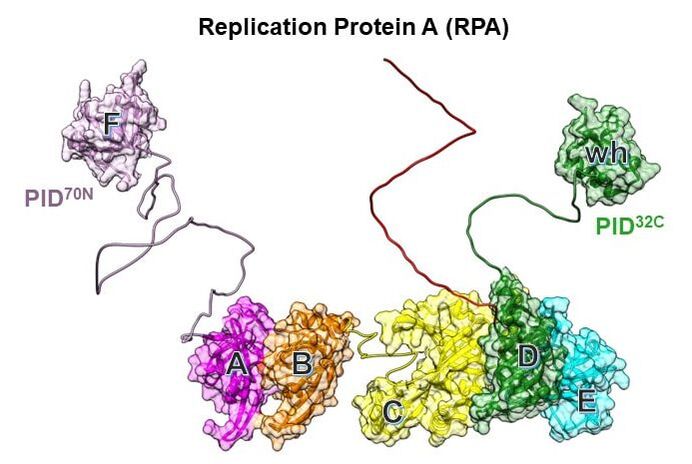
Replication Protein A and the coordination of DNA metabolism
DNA metabolic processes including replication, repair, recombination, and telomere maintenance occur on single-stranded DNA (ssDNA). In each of these complex processes, dozens of proteins function together on the ssDNA template. However, when double-stranded DNA is unwound, the transiently open ssDNA is protected and coated by the high affinity heterotrimeric ssDNA binding Replication Protein A (RPA). Almost all downstream DNA processes must first remodel/remove RPA or function alongside to access the ssDNA occluded under RPA. Formation of RPA-ssDNA complexes trigger the DNA damage checkpoint response and is a key step in activating most DNA repair and recombination pathways. Thus, in addition to protecting the exposed ssDNA, RPA functions as a gatekeeper to define functional specificity in DNA maintenance and genomic integrity. The precise mechanisms of how RPA imparts functional specificity is poorly resolved. Our long-term goals are to answer the following questions: RPA physically interacts with over three dozen DNA processing enzymes. How are these interactions determined, regulated, and prioritized? RPA binds to ssDNA with high affinity. How do DNA metabolic enzymes that bind to ssDNA with hundred-fold lower affinities remove RPA? RPA plays a role in positioning the recruited enzymes (with appropriate polarity) onto the DNA. What are the structural, kinetic, and thermodynamic properties that regulate this process? How are the DNA and protein interaction activities of RPA tuned by post translational modifications such as phosphorylation? Utilizing our powerful biochemical, structural, and biophysical toolkit we seek to resolve how RPA functions coordinates the many DNA metabolic processes.
Homologous recombination and
maintenance of genome integrity and stability
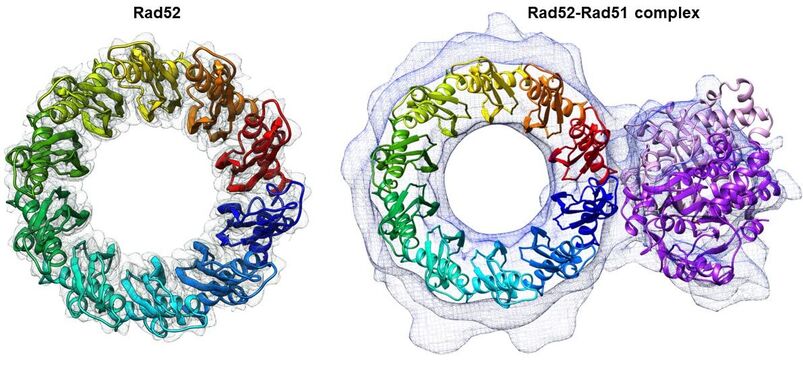
Double-stranded DNA breaks (DSBs) are drivers of genomic instability and associated cancers. On average, ~10 to 50 DSBs occur per cell per day and thus pose the highest risk to genomic integrity. Homologous Recombination (HR), single strand annealing (SSA), and RNA-templated DNA repair (RDR) are key biological processes that protect the cell from DSBs. The fundamental problem with a DSB is that physical continuity is broken within a chromosome. Thus, for accurate repair, the undamaged sister allele is used as a template to copy genetic information. Homology directed DSB repair transpires through either Rad51-dependent HR or Rad51-independent SSA pathways. For both processes, the DSB must first be processed (resected) to generate 3′ single strand DNA (ssDNA) tails which are coated by the ssDNA binding Replication Protein A (RPA). Binding of RPA triggers activation of the ATR kinase and associated repair signaling mechanisms. In HR, mediator proteins such as Rad52 and BRCA2 promote loading of the Rad51 protein (recombinase) onto RPA-coated ssDNA. The Rad51-coated 3′-ssDNA filament invades the undamaged sister chromatid and base pairs with the complementary region, which then is used as a template to replicate the missing information. In certain cases, when the 3′-ssDNA overhangs contain highly homologous direct repeats, or during replication stress in common fragile sites, SSA is triggered. Here, Rad52 promotes the direct annealing of the direct repeats between the two 3′-ssDNA overhangs. The non-annealed 3′-flaps are excised, and the resulting structures are subsequently processed by DNA repair synthesis. While HR is an accurate process for DSB repair, SSA is relatively error prone. Finally, RNA can serve as a template for DSB repair where R-loop intermediates (RNA-DNA hybrids) can be used as substrate to copy the missing information. Our work focuses on how pro- and anti-HR mediator proteins regulate formation of the Rad51 filament and subsequent downstream events. Current focus is on the mechanism of action of Rad52 (a pro HR mediator) and Srs2 (an anti-HR mediator). Utilizing biophysical, mass-spectrometry, single-molecule fluorescence, and cryoEM approaches we seek to uncover how these enzymes coordinate homologous recombination.
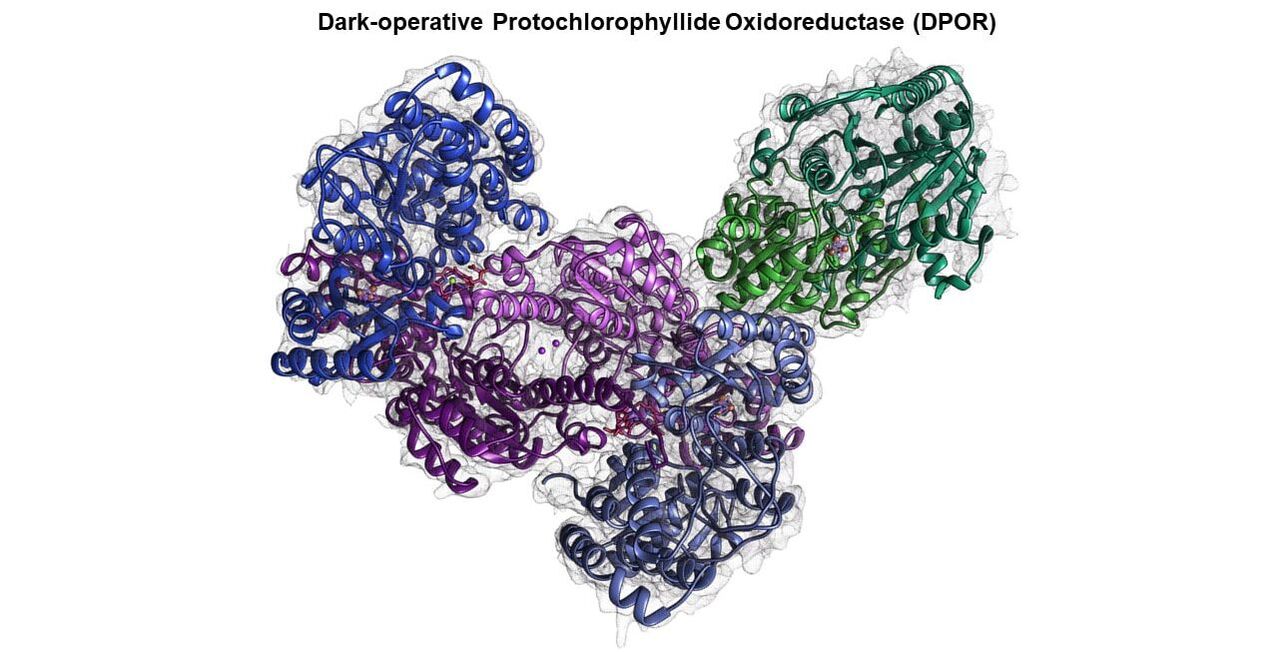
Mechanisms of long-range electron transfer in oxidoreductases
Our long-term goals are to understand how nitrogenase and nitrogenase-like enzymes catalyze ATP-dependent multi-electron substrate reduction reactions. The architecture of these enzymes and how they mediate long-range electron transfer (ET) over a series of sophisticated metal centers have served as blueprints for the design of bio-inspired catalysts. Nitrogenase fixes atmospheric dinitrogen in the biological nitrogen cycle, and both DPOR (Dark-operative Protochlorophyllide Oxidoreductase) and COR (Chlorophyllide a Oxidoreductase) catalyze key reduction reactions in the ‘dark’ chlorophyll biosynthetic pathway and are essential for photosynthesis. In addition to their significance in life-sustaining fundamental biological processes, DPOR, COR, and nitrogenase serve as excellent model systems to decipher complex long-range electron transfer processes in biology. These enzymes are made of homodimeric electron donor and heterotetrameric electron acceptor component proteins that transiently assemble in the presence of ATP to perform electron transfer. The heterotetrameric acceptor component possesses two identical active sites. We uncovered that the two halves in nitrogenase and DPOR function in an asymmetric manner with catalytic events in one half allosterically controlling the other. Perturbing activities in one half stalls substrate reduction in the complex. The functional significance of this asymmetry and the mechanistic basis of allosteric control are not understood. We seek to decipher how assembly of the component proteins, electron transfer, ATP binding and hydrolysis, and substrate reduction contribute to establishing and coordinating allostery within the enzyme complex. These studies will help in our ability to design artificial catalysts for difficult reduction chemistries.
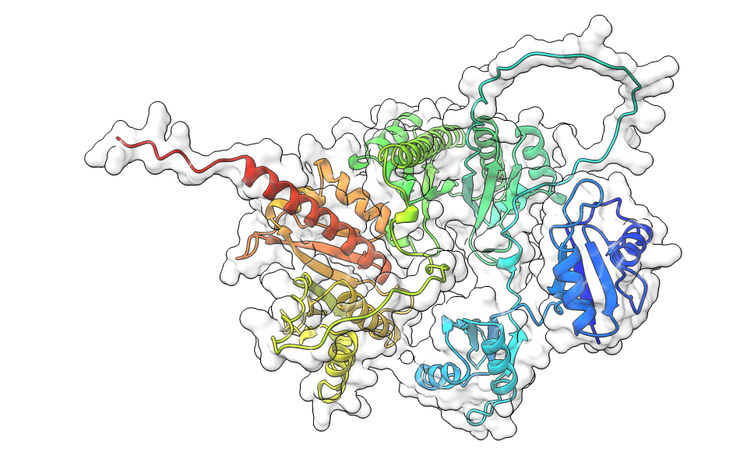
mRNA recognition by the oncofetal IMP family of RNA binding proteins
RNA binding proteins recognize hundreds of mRNA targets and regulate their expression through splicing, end processing, nuclear export, translational control, and altering mRNA stability. Most such proteins lack enzymatic activity but serve as a scaffold for the recruitment of other proteins onto the mRNA. The assembly of such complexes, on the appropriate mRNA target, confer specificity to accurate and rapid gene expression demands within the cell. A small number of RNA binding proteins control the expression of thousands of mRNAs, and our long-term goal is to decipher how these proteins gain specificity for a defined mRNA target. In other words, how do these proteins pick out one mRNA from thousands of potential mRNA substrates? We address this question in the IMP family of mRNA binding proteins. Insulin-like growth factor 2 mRNA binding proteins (IMPs) is a family of three paralogs (IMP1, IMP2 and IMP3) and functions in normal development, stem cell maintenance, and when deregulated leads to cancers. These proteins use six high affinity RNA binding domains tethered by flexible linkers, thus making them challenging to investigate through traditional biochemical approaches. Our work aims to: 1) Obtain structural and mechanistic knowledge of how these enzymes function. 2) Probe how each member contributes to embryonic development. 3) Why do mutations and re-expression lead to tumor development.
Join our team
We welcome applications from undergraduate students, graduate students, postdoctoral scholars, and research scientists to be a part of our team and explore these enzymes. We provide a collaborative research environment well supported by extramural and internal funding. We take pride in our attention to your personal development and career needs, and is a central pillar that supports all our science. We also welcome middle school and high school students seeking to perform research in the summers.
Please browse through our content and feel free to reach out: CONTACT
Please browse through our content and feel free to reach out: CONTACT
Our research is supported by grants from the following federal and private agencies